 For the majority of the last 100 years the role of bacteria in human health has been explored in terms of risk to health and well-being, the ‘bad bug = bad health’ paradigm. The result has been a combination of remarkable benefits against infectious related deaths and a slow but steady development of chronic non communicable diseases (CNCDs) – cardiovascular disease, cancer, diabetes and respiratory diseases now kill more people worldwide than all other diseases combined.
For the majority of the last 100 years the role of bacteria in human health has been explored in terms of risk to health and well-being, the ‘bad bug = bad health’ paradigm. The result has been a combination of remarkable benefits against infectious related deaths and a slow but steady development of chronic non communicable diseases (CNCDs) – cardiovascular disease, cancer, diabetes and respiratory diseases now kill more people worldwide than all other diseases combined.
This rate of demise will continue to rise in the coming years as the global population ages, sedentary lifestyles and inappropriate food consumption continues to spread across the world.
In the world of nutritional and functional medicine the possibilities of harnessing rather than destroying the bacteria we share our moist tissues with has existed in a evolved manner since Élie Metchnikoff, (Mechnikoff’s work on phagocytes won him the Nobel Prize in 1908) proposed the concept of bacterial therapy through the consumption of fermented foods as a mechanism of delivering health benefits to the consumer from the end of the 19th Century.
So am I simply about to repeat much of what has gone before in this short piece, or is there something else we need to consider?

Intersecting disciplines
Without doubt, the intersecting fields of immunology, microbiology, nutrition and medicine are fast evolving, driven by extensive investment in research, such as the microbiome project.[1] However, as a result of all this new data, many researchers are now asking potentially awkward questions about the nature and validity of current and historical immune research in light of our greater understanding of the impact of commensal bacteria on lab and life outcomes.[2]
Has the scientific community been labouring away without realising that despite their carefully controlled lab conditions, and genetically identical animal models, that their experimental charges – the world of mice and rats as well as humans have been producing different outcomes because of the bacterial mixes they harbour in their guts?
Research and Bugs
The relatively recent recognition that certain specific bacteria can influence the formation of important sub sets of our primary defence cells; called T cells, has generated a great deal of clinical interest in the immune enhancing ability of the bacteria we harbour in our digestives tract.[3] The bacteria and their environmental triggers have the consequence of imparting soluble signalling chemicals that change the phenotype (the form) of effector T cell available for our use in both acute and chronic health management.
Of greater interest to clinicians is the discovery that these peripherally induced phenotypes are reversible, offering opportunities for recovery, and for loss of benefit if environmental triggers, such as probiotics, micronutrients and diet are discontinued too soon. Whilst these individual cells may seem quite complex for most practitioners to learn about, the regulatory T cells and effector (cells that impart an effect on other cells) Tcells such as Th17 have important implications for immunologists and ultimately practitioners as they are significant factors that drive molecular inflammation, the ‘sine qua non’ of non infectious chronic illness.[4],[5]
The diagram below shows in a schematic the interactions we should be considering between our own DNA and those of our microbiota. Simplifies to a cartoon, it illustrates that we need to consider ourselves as a super organism, made up of both human and bacterial DNA, and actually outnumbered in terms of genetic material, by 100 times our bacterial hitchhikers. The idea that only increased pro-inflammatory bacteria have a deleterious effect on T cell balance is also challenged, as too few anti-inflammatory organisms will also allow development of para-inflammation, a persistent low grade inflammation capable of inducing illness.[6]
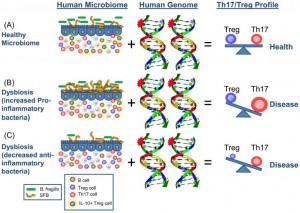
The results of this symbiotic relationship between food, bacteria and us, are not simply confined to the digestive tract. It is apparent that their combined spread their effects beyond the local tissues and influence a wide range of non-infectious and infectious chronic disorders. These include allergies, asthma, diabetes, obesity, cancer and, quite remarkably, very recently identified neuropathology’s.[8]
Impact on Research
This then means that researchers studying the immune system that exclude the effect of their experimental charges bacterial composition on challenge and outcome may be missing important variables out of their data. Even small changes in the bacterial population can have a significant impact on outcome.[9]
As I have previously discussed, faecal transplantation and antibiotic usage have quite significant impact on bacterial composition, and not only for a few days, as was strongly proposed for many years, but in some cases months or even permanently.[10] It has long been understood that bacteria in the gut have a significant impact on the metabolism of drugs which in turn influence disease outcomes and should be taken in context with individual genetic traits as well.[11] We need to keep in mind that these bacteria are also likely to influence differing outcomes via the metabolism of food supplements, many of which rely of liver deamination and bacterial degradation prior to cellular uptake.
As one might expect, the choice of diet and environmental differences can have an impact on human and animal bacterial populations, even in otherwise genetically identical mice. This effects the growth and development of the bacteria leading to different intestinal microbiota populations. Even modest stress and a change of room can have relevant effects on the bacterial compositions leading to different outcomes.[12],[13]
These are just some examples that highlight the significant impact that the microbiota has in experimental models involving various aspects of the host immune response and should indicate to all of us that if we are to pursue the idea of personalised health care that the genomic traits in our own DNA may be less valid in terms of treatment options than the genetic variations in our commensal bed fellows.
Future Clinical Investigations.
How might this be employed in a clinical setting, as collecting the microbial DNA is well beyond the clinical remit of most practitioners and even collecting a stool sample for broad range analysis presents many problems, not least relevance and validity issues. Well the answer may lie in the collection of urinary metabolites, which have already been shown to vary based on alterations to the microbial community.[14] But whilst this is certainly an attractive option there is no standardisation of metabolites against relevant microbial balances and changes.
We are witnessing a sea change in researchers understanding of the role of microbial organisms beyond those attributed via Koch’s postulates to a pathogenic role. As practitioners well versed in the concepts of adding probiotics; a temporary community of hopefully advantageous organisms. It may be time for us to start to ask more demanding questions of our bacterial manufacturers, such as what immunological effects are engendered in a cross section of the population when we consume these innocuous bacteria, and in doing so, may we inadvertently reduce the impact of other agents rather than amplifying them.
I have long discussed the need for strain specific probiotic research and the main research centres involved in bacterial analysis are increasingly relying on dedicated characterisation of species to accurately reflect changes in the individual’s metabolism and immune response.[15] Perhaps this an indication of a drawing to a close the traditional ecological view – just take probiotics – they can only do you good, and refining the future strategy to: take these probiotics as they meet your immunological phenotypical needs!
References
[1] http://commonfund.nih.gov/hmp/ [visited 18th Oct 2011]
[2] Lee, Y. K. & Mazmanian, S. K. Has the microbiota played a critical role in the evolution of the adaptive immune system? Science 330, 1768–1773 (2010). View Abstract
[3] Jarchum I, Pamer EG. Regulation of innate and adaptive immunity by the commensal microbiota. Curr Opin Immunol. 2011 Jun;23(3):353-60. Epub 2011 Apr 3. Review. View Abstract
[4] Feng, T. & Elson, C. O. Adaptive immunity in the host–microbiota dialog. Mucosal Immunol. 4, 15–21 (2011). View Abstract
[5] Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341 (2011). View Abstract
[6] Medzhitov R. Inflammation 2010: new adventures of an old flame. Cell. 2010 Mar 19;140(6):771-6. View Abstract
[7] Lee YK, Mazmanian SK. Has the microbiota played a critical role in the evolution of the adaptive immune system? Science. 2010 Dec 24;330(6012):1768-73. Review. View Abstract
[8] Bischoff, S. C. ‘Gut health’: a new objective in medicine? BMC Med. 9, 24 (2011). View Full Paper
[9] Sekirov, I. et al. Antibiotic-induced perturbations of the intestinal microbiota alter host susceptibility to enteric infection. Infect. Immun. 76, 4726–4736 (2008). View Abstract
[10] Wlodarska M, & Finlay BB (2010). Host immune response to antibiotic perturbation of the microbiota. Mucosal immunology, 3 (2), 100-3 PMID: View Abstract
[11] Jia, W., Li, H., Zhao, L. & Nicholson, J. K. Gut microbiota: a potential new territory for drug targeting. Nature Rev. Drug Discov. 7, 123–129 (2008). View Abstract
[12] Friswell, M. K. et al. Site and strain-specific variation in gut microbiota profiles and metabolism in experimental mice. PLoS ONE 5, e8584 (2010). View Abstract
[13] Bailey, M. T. et al. Stressor exposure disrupts commensal microbial populations in the intestines and leads to increased colonization by Citrobacter rodentium. Infect. Immun. 78, 1509–1519 (2010). View Abstract
[14] Zheng X, Xie G, Zhao A, Zhao L, Yao C, Chiu NH, Zhou Z, Bao Y, Jia W, Nicholson JK, Jia W. The Footprints of Gut Microbial-Mammalian Co-Metabolism. J Proteome Res. 2011 Oct 4 View Abstract
[15] Quigley EM. Gut microbiota and the role of probiotics in therapy. Curr Opin Pharmacol. 2011 Oct 11 View Abstract





